
Open-Source Internship opportunity by OpenGenus for programmers. Apply now.
In this article, we will cover 2 different convolution methods: Kn2row and Kn2col Convolution which are alternatives to Im2row and Im2col. Kn2row and Kn2col are space efficient variants.
Table of Contents:
- Introduction.
- Kn2row and Kn2col Convolution
2.1 Kn2row Algorithm
2.2 Kn2col Algorithm - Differences.
- Memory Usage.
- GEMM Call for Kn2row.
- GEMM call for Kn2col.
- Implementation of kn2row convolution.
- Implementation of kn2col convolution.
- Quick Summary of all types of Convolution methods.
Pre-requisites:
Introduction:
Convolution is the basic process of applying a filter to an input to produce an activation. The position and strength of a recognised feature in an input, such as an image, are shown by a feature map created by repeatedly applying the same filter to the input.
The capacity of convolutional neural networks to learn a large number of filters in parallel particular to a training dataset under the restrictions of a given predictive modelling task, such as image classification, is what makes them unique. As a result, extremely particular characteristics may be recognised across the input photos.
Kn2row and Kn2col Convolution:
Kn2row: Also Known as kernel to row, This approach is based on the fact that a KxK convolution may be computed by first shifting and combining the partial outputs from K.K 1x1 convolutions. MxHxW is the additional buffer space necessary for this.
Kn2col: Also known as kernel to Column is the inverse of the kn2row approach.
One thing we know that, we can compute 1 × 1 MCMK without data replication, but how can one implement K × K MCMK, for K > 1?
A K ×K convolution can be expressed as the sum of K^2(k square) separate 1 × 1 convolutions. The sum, on the other hand, is not easy to calculate. Each one-to-one convolution produces a result matrix with the dimensions [M]x [H x W]. Because each resultant matrix corresponds to a different kernel value in the K x K kernel, we can't just add each of the resulting matrices pointwise. The addition of these matrices can then be resolved by vertically and/or horizontally (row and column offsets) offsetting every pixel in every channel of the multi-channel representation of these matrices by one or more places before the addition.
For example, when computing a 3×3 convolution the result from computing the 1 × 1 MCMK for the central point of the 3 × 3 kernel is perfectly aligned with the final sum matrix. On the other hand, the matrix that results from computing the 1×1 MCMK for the upper left value of the 3×3 kernel must be offset up by one place and left by one place (in its multichannel representation) before being added to the final sum that computes the 3 × 3 MCMK. Note that when intermediate results of 1 × 1 convolutions are offset, some values of the offsetted matrix fall outside the boundaries of the final result matrix. These out-of-bounds values are simply discarded when computing the sum of 1 × 1 convolutions.
Let’s see the Kn2row Algorithm:
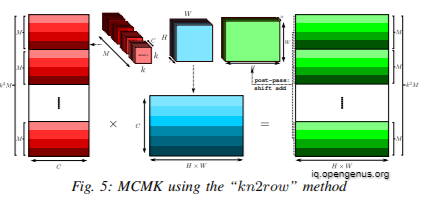
A single matrix multiplication may be used to compute each of the K^2 separate 1 x 1 convolutions. We reorganise the kernel matrix such that the channel data is set out in a logical manner, with M being the outer dimension and C being the inner dimension. This data reorganisation may be done statically ahead of time and used for all subsequent MCMK invocations. We multiply a [K^2xM]x[C] kernel matrix by a [C]x[HxW] input matrix with a single call to GEMM, resulting in a [K^2xM]X[HXW] matrix. In the multi-channel representation, we do a post pass of shift-add by summing each of the M^2 submatrices of size M x [H x W] with proper offsets. The output of our MCMK method is a [M] x [H x W] matrix, which is the result of this sum. This is the kn2row algorithm.
Let’s see the Kn2col Algorithm:
Inorder to get the kn2col Algorithm we have to swap the dimensions of the kernel matrix so that C is not the innermost dimension and swap the input layout to make C the innermost dimension, resulting in the kn2col algorithm.
Difference:
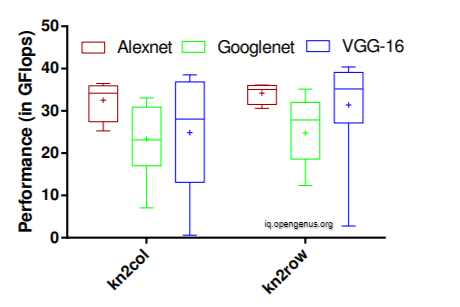
The kn2row approach has a higher degree of localisation. In all of the networks, the kn2row creates a significantly narrower dispersion of performance between layers, most notably in AlexNet. Another noteworthy finding is that the performance of both kn2col and kn2row increases with C and M while decreasing with kernel size.
Memory Usage:
When we will evaluate our algorithm, we may find out that for a number of our chosen input dimensions the board ran out of memory intermittetly while executing our O(K^2) algorithms. Sometimes they require extra memory and with respect to our topic, the methods take up extra space as follows:
Kn2row:
(K^2 − 1) × (M × (HW))
Kn2col:
Kn2col: (K^2 − 1) × (M × (HW))
Now let's look at the General Matrix Multiplication of the both the convolution methods.
Gemm Call for Kn2row:
[K^2×M]×[C] // kernel matrix
[C]×[H×W] // input matrix
We multiply kernel matrix by input matrix
[K^2×M]×[C] x [C]×[H×W]
= [K2×M]×[H×W]
perform a post pass of shift-add by summing each of the M^2 submatrices.
= [M] × [H × W] // output of our MCMK algorithm
Gemm Call for Kn2col:
[H×W]× [C] // input matrix
[C]×[K2 ×M] // kernel matrix
We multiply input matrix by kernel matrix:
[H×W]× [C] x [C]×[K2 ×M]
=[H × W] × [K2 × M] // matrix
Implementation for the Kn2row Convolution method:
We will have a look at the implementation of kn2row convolution in C
#include <assert.h>
#include <stdbool.h>
#include <stdlib.h>
#include <string.h>
#include <cblas.h>
#include "common_types.h"
#include "data_reshape.h"
#include "utils.h"
//
// col_shift : +ve --> shift left overlap mat , -ve --> shift right overlap mat
// or shift left base mat and keep overlap mat as it is.
//
//
// row_shift : +ve (coeff is down the center coeff) --> shift up overlap mat ,
// -ve --> shift down overlap mat or shift up the base mat.
void MatrixShiftAdd(float *base_mat,
int base_no_rows, int base_no_cols,
float *overlap_mat,
int ov_no_rows, int ov_no_cols,
int row_shift, int col_shift) {
if (row_shift == 0 && col_shift == 0 && (base_no_rows == ov_no_rows) &&
(base_no_cols == ov_no_cols)) {
// normal matrix add
cblas_saxpy(base_no_rows * base_no_cols, 1.0, overlap_mat, 1, base_mat, 1);
return;
}
int rows_to_add, cols_to_add;
int base_row_start, base_col_start;
int ov_row_start, ov_col_start;
// without padding case
if (ov_no_rows > base_no_rows) {
rows_to_add = base_no_rows;
cols_to_add = base_no_cols;
base_row_start = 0;
base_col_start = 0;
ov_row_start = row_shift < 0? -row_shift : 0;
ov_col_start = col_shift < 0? -col_shift : 0;
} else {
rows_to_add = ov_no_rows - abs(row_shift);
cols_to_add = ov_no_cols - abs(col_shift);
ov_col_start = col_shift > 0? col_shift : 0;
ov_row_start = row_shift > 0? row_shift : 0;
base_row_start = row_shift < 0? -row_shift : 0;
base_col_start = col_shift < 0? -col_shift : 0;
}
for (int r = 0; r < rows_to_add; ++r) {
int base_mat_offset = (r + base_row_start) * base_no_cols + base_col_start;
int overlap_mat_offset = (r + ov_row_start) * ov_no_cols + ov_col_start;
cblas_saxpy(cols_to_add, 1.0, overlap_mat + overlap_mat_offset, 1,
base_mat + base_mat_offset, 1);
}
}
bool Kn2RowConvLayer(const float *in_data, const float *filters,
const float *bias, TensorDim in_dim,
TensorDim filt_dim, int stride, int pad, int group,
float *output) {
assert(group == 1);
assert((pad == 0) || (pad == filt_dim.w / 2));
assert(in_dim.n == 1);
assert(filt_dim.h == filt_dim.w);
assert(stride == 1);
// Output dimensions.
TensorDim out_dim;
out_dim.w = (in_dim.w + (pad + pad) - filt_dim.w) / stride + 1;
out_dim.h = (in_dim.h + (pad + pad) - filt_dim.h) / stride + 1;
out_dim.c = filt_dim.n;
out_dim.n = in_dim.n;
// Re-arrange filters in the k x k x no_out_maps x no_in_maps.
// We can avoid this if the filters are already reshaped in this format.
float *kkmc_filters = malloc(filt_dim.n * filt_dim.c * filt_dim.h *
filt_dim.w * sizeof(float));
NCHW2HWNC(filters, filt_dim.n, filt_dim.c, filt_dim.h, filt_dim.w,
kkmc_filters);
int H = in_dim.h;
int W = in_dim.w;
float alpha = 1.0;
float beta = 0.0;
// We need separate buffer because GEMM output will have width = H*W even
// if there is no padding (pad = 0).
float *gemm_output = malloc(out_dim.c * H * W * sizeof(float));
// Prefill output buffer with bias if present else set to zero.
if (bias) {
for (int m = 0; m < out_dim.c; ++m) {
for (int a = 0; a < out_dim.h * out_dim.w; ++a) {
output[m * out_dim.h * out_dim.w + a] = bias[m];
}
// For batch size > 1
for (int b = 1; b < out_dim.n; ++b) {
memcpy(output + b * out_dim.c * out_dim.h * out_dim.w,
output, out_dim.c * out_dim.h * out_dim.w * sizeof(float));
}
}
} else {
memset(output, 0, out_dim.n * out_dim.c * out_dim.h * out_dim.w *
sizeof(float));
}
for (int kr = 0; kr < filt_dim.h; kr++) {
int row_shift = kr - filt_dim.h / 2;
for (int kc = 0; kc < filt_dim.w; kc++) {
int group_no = kr * filt_dim.w + kc;
int col_shift = kc - filt_dim.w / 2;
// Matrix dimensions - A -> mxk B -> kxn C --> mxn
int m = filt_dim.n;
int k = filt_dim.c;
int n = in_dim.h * in_dim.w;
// This is just 1x1 convolution
cblas_sgemm(CblasRowMajor, CblasNoTrans, CblasNoTrans,
m, n, k, alpha, kkmc_filters + group_no * m * k,
k, in_data, n, beta, gemm_output, n);
// Slide the resulting matrix which has contribution from one of the
// KxK kernel coefficients and add to the output.
for (int omap = 0; omap < filt_dim.n; omap++) {
MatrixShiftAdd(output + omap * out_dim.h * out_dim.w,
out_dim.h, out_dim.w,
gemm_output + omap * H * W,
H, W, row_shift, col_shift);
}
}
}
free(kkmc_filters);
free(gemm_output);
return true;
}
Implementation of Kn2col convolution methods:
We will have a look at the implementation of kn2col convolution in C:
#include <stdbool.h>
#include <cblas.h>
#include <assert.h>
#include <stdlib.h>
#include <string.h>
#include "common_types.h"
#include "data_reshape.h"
#include "utils.h"
#include "conv_layers.h"
bool Kn2ColConvLayer(const float *in_data, const float *filters,
const float *bias, TensorDim in_dim,
TensorDim filt_dim, int stride, int pad, int group,
float *output) {
assert(group == 1);
assert((pad == 0) || (pad == filt_dim.w / 2));
assert(in_dim.n == 1);
assert(filt_dim.h == filt_dim.w);
assert(stride == 1);
// Output dimensions.
TensorDim out_dim;
out_dim.w = (in_dim.w + (pad + pad) - filt_dim.w) / stride + 1;
out_dim.h = (in_dim.h + (pad + pad) - filt_dim.h) / stride + 1;
out_dim.c = filt_dim.n;
out_dim.n = in_dim.n;
// Reshape filters in CHWN (ker_size x ker_size x no_in_ch x no_out_ch) format
float *kkcm_filters = malloc(filt_dim.n * filt_dim.c * filt_dim.h *
filt_dim.w * sizeof(float));
NCHW2HWCN(filters, filt_dim.n, filt_dim.c, filt_dim.h, filt_dim.w,
kkcm_filters);
int H = in_dim.h;
int W = in_dim.w;
float alpha = 1.0;
float beta = 0.0;
// We need separate buffer because GEMM output will have width = H*W even
// if there is no padding (pad = 0).
float *gemm_output = malloc(out_dim.c * H * W * sizeof(float));
float *nchw_gemm_output = malloc(out_dim.c * H * W * sizeof(float));
// Prefill output buffer with bias if present else set to zero.
if (bias) {
for (int m = 0; m < out_dim.c; ++m) {
for (int a = 0; a < out_dim.h * out_dim.w; ++a) {
output[m * out_dim.h * out_dim.w + a] = bias[m];
}
// For batch size > 1
for (int b = 1; b < out_dim.n; ++b) {
memcpy(output + b * out_dim.c * out_dim.h * out_dim.w,
output, out_dim.c * out_dim.h * out_dim.w * sizeof(float));
}
}
} else {
memset(output, 0, out_dim.n * out_dim.c * out_dim.h * out_dim.w *
sizeof(float));
}
for (int kr = 0; kr < filt_dim.h; kr++) {
int row_shift = pad - kr;
for (int kc = 0; kc < filt_dim.w; kc++) {
int group_no = kr * filt_dim.w + kc;
int col_shift = pad - kc;
// Matrix dimensions - A -> mxk B -> kxn C --> mxn
int m = in_dim.h * in_dim.w;
int k = filt_dim.c;
int n = filt_dim.n;
// output is in H x W x C format.
cblas_sgemm(CblasRowMajor, CblasTrans, CblasNoTrans,
m, n, k, alpha, in_data, m,
kkcm_filters + group_no * filt_dim.c * filt_dim.n, n, beta,
gemm_output, n);
// convert to CxHxW format.
// FIXME: this will be slow. Need to find other ways :(
NHWC2NCHW(gemm_output, 1, filt_dim.n, H, W, nchw_gemm_output);
for (int omap = 0; omap < filt_dim.n; omap++) {
MatrixShiftAdd(output + omap * out_dim.h * out_dim.w,
out_dim.h, out_dim.w,
nchw_gemm_output + omap * H * W,
H, W, row_shift, col_shift);
}
}
}
free(kkcm_filters);
free(gemm_output);
free(nchw_gemm_output);
return true;
}
Please note the following notations:
N : Batch size
C : No of input channels
H : Input height
W : Input width
M : No of output channels
K : Kernel size
We at Opengenus have covered almost all the major types of Convolution i.e. Im2row, Im2col, Kn2row and Kn2col, here is a quick definition and benefit of every convolution method:
Im2row vs Im2col:
1. Im2col: Im2col convolution, also known as Image Block to Column, is a method that involves flattening each window and stacking it as columns in a matrix.
2. Im2row : Im2row is a method wherein the local patches are unrolled into rows of the input-patch matrix I and the kernels are unrolled into columns of the kernel-patch-matrix K. Im2row works better compared to im2col.
Comparison/Benefit:
Im2col:The memory footprint of the input matrix is considerably increased by this im2col conversion, which lowers data locality.
Im2row: Im2row Convolution is actually a better version of im2col convolution. It is fast and steady and improves model performance.
Kn2row vs Kn2col:
1. Kn2row: This approach is based on the fact that a KxK convolution may be computed by first shifting and combining the partial outputs from K.K 1x1 convolutions. MxHxW is the additional buffer space necessary for this.
2. Kn2col: Also known as kernel to Column this method is the exact inverse of kn2row convolution method.
Comparison/Benefit:
The kn2row also produces a slightly tighter spread of performance across layers in all the networks, most notably in AlexNet.
However, kn2col and kn2row both tend to increase with the C and M and decreases with the size of the kernel.
With this article at OpenGenus, you must have the complete idea of kn2row and kn2col Convolution.
