
Open-Source Internship opportunity by OpenGenus for programmers. Apply now.
Medical image processing is an important application in Computer Vision,requires segmentation of images into body parts. Joseph Bullock and his partners in Durham University proposed a neuron network called XNet which is suitable for this task. Only trained on a small dataset,XNet still surpassed classical methods to achieve state-of-the art result.In this article,I'll cover main points of the paper.
Related work
Before exploring content of this paper, we need to go through some groundworks which XNet's method based on.Previously,much work focused on using classical image processing techniques,especially pixel clustering based on similarity in certain parameters.The authors dived into those methods and drew crucial insights in table 1:
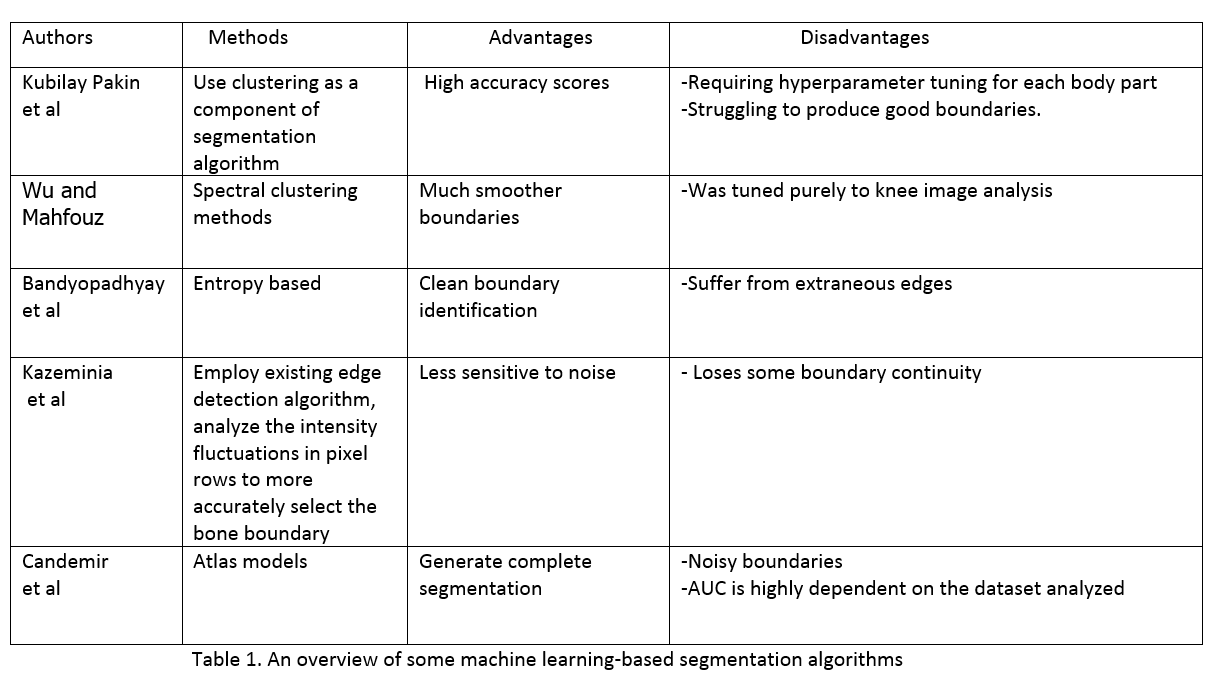
Overall, classical techniques are not good enough and suitable for wide application in X-ray analising due to their architectures.Therefore, Joseph Bullock and his parters pay their attention to neuron network-based methods which currently perform state-of-the-art result and can meet their requirements.This study was based on U-Net and Segnet model which show good performance and widely applied.However,some variants of them have been largely constrained to the segmentation of cell structures or only used in specific cases.
Data
The researchers collected 150 images which are composed of 69 CT scans images of feet,knees and phantom head, and 81 standard X-Ray images of different body and phantom body parts, of which the thorax is the most underrepresented.These images were pre-processed by utilize mean-subtraction and pixel value normalisation to within the range [−1, 1] and also resized to 200x200 pixels to match the input shape of network.There are three distinct regions in every iamge: open beam, bone and soft tissue.Joseph Bullock and his parters labelled images using free software GIMP by assigning each color to one of these three regions:yellow to bone,green to soft tissue and purple to soft tissue.See the figure 3 of the paper:
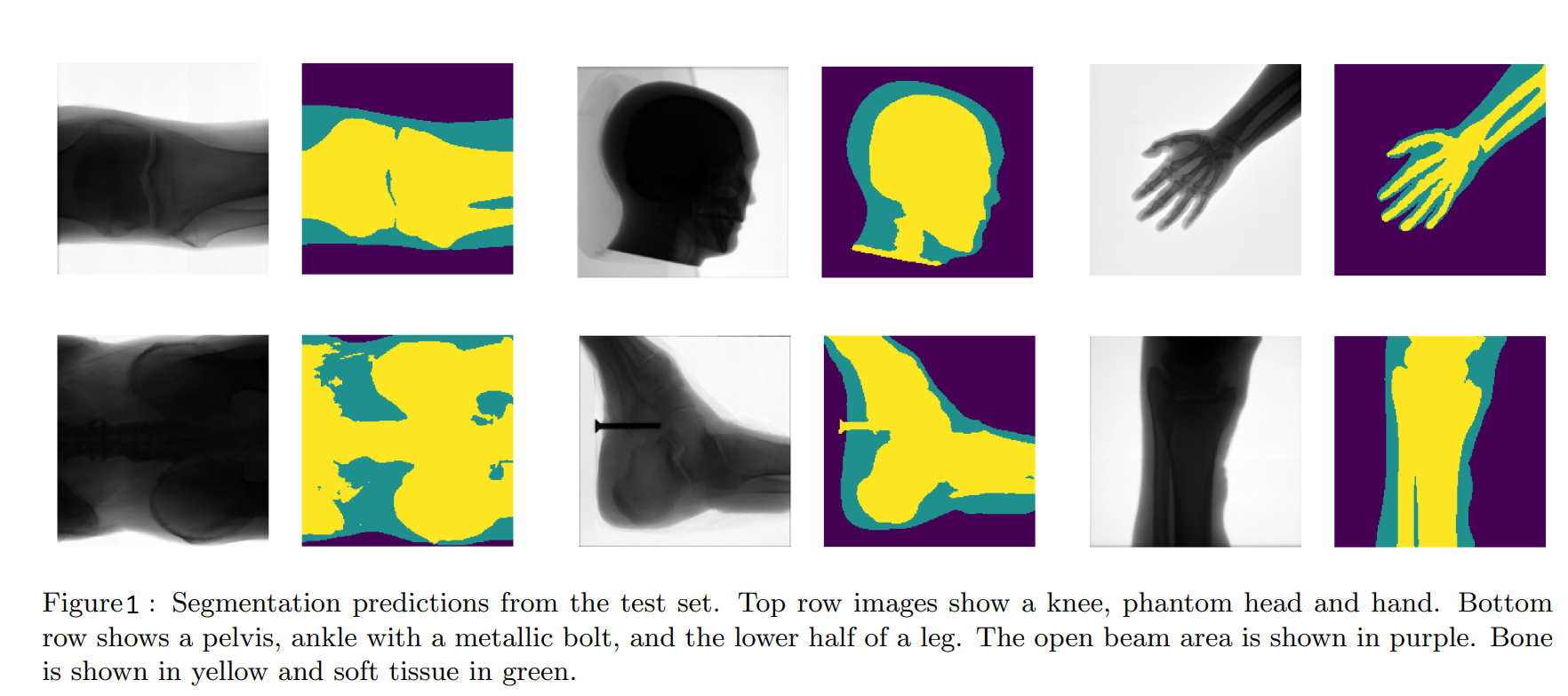
Note that several images contain foreign metal objects and we classify them as bone.
There are 19 different body parts and their number of images are shown in the table 2.
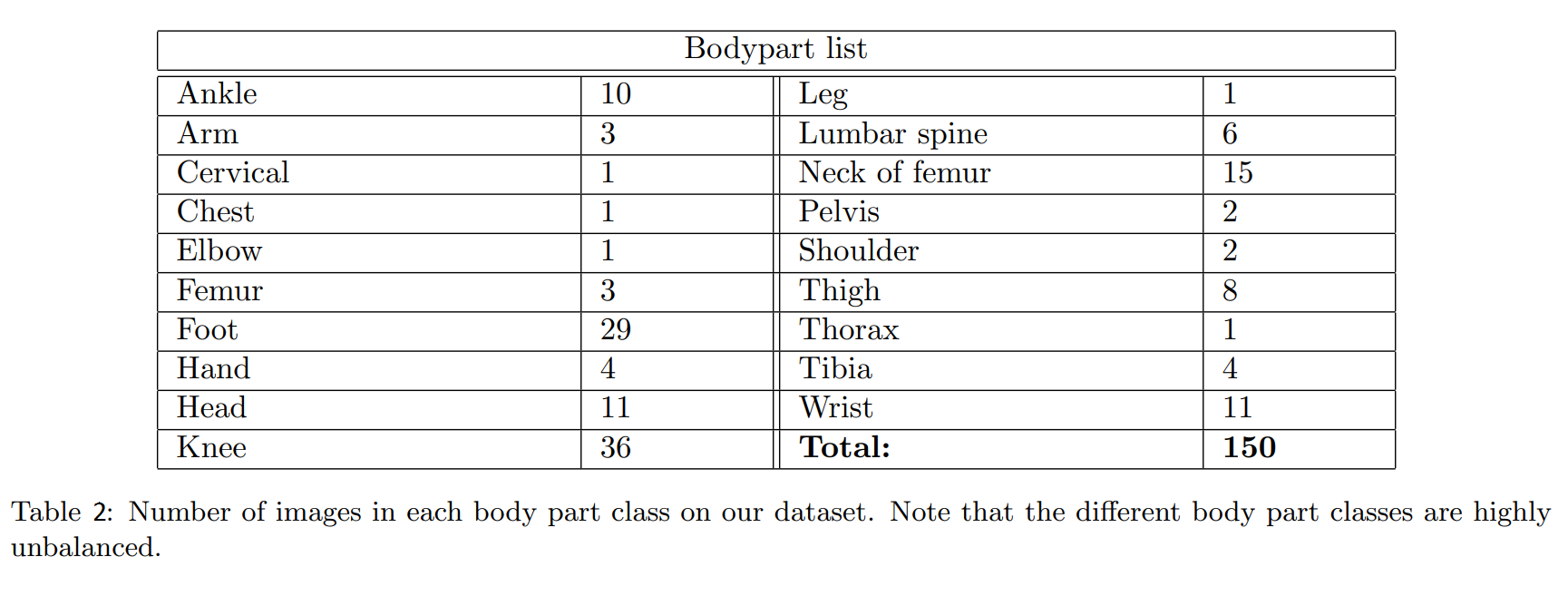
It is clear that our dataset is too small and imbalanced.If we use this dataset for complex model like XNet, it's probably overfitting.In order to avoid that, the researchers augmented data. After experimenting with a variety of filters,they chose elastic transformation because it's important for generating realistic augmented images,combined with other transformations like translation,cropping,rotation,shear.
This technique produced 500 images per body part class.So, to conclude, the authors augmented data for 2 purposes: avoiding overfitting and balancing the different body part classes through oversampling.
Architecture
The structure of XNet consists of 2 parts:Encoder and Decoder.See the figure below:
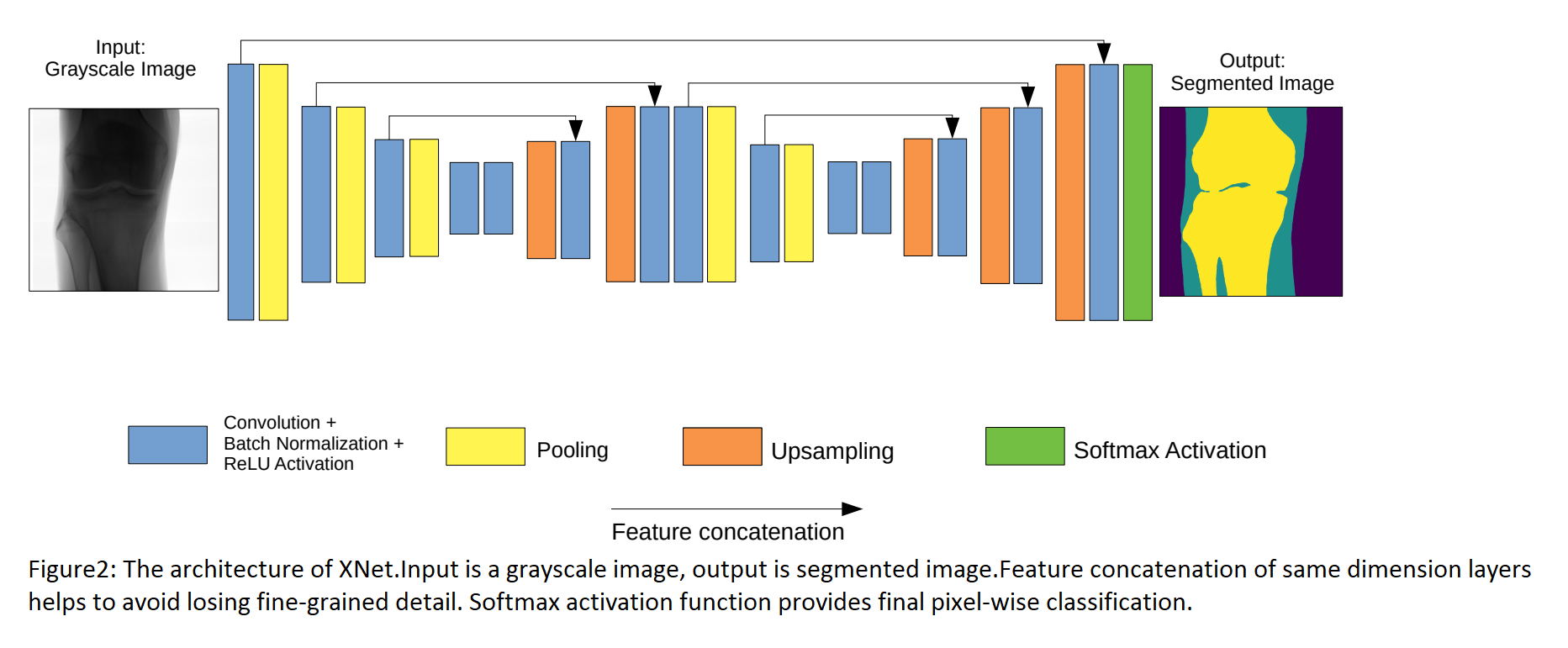
Encoder
This component lies mainly on the left of the picture.It is composed of many convolutional layers and pooling layers.After each convolutional layer, the number of feature maps increases but their size decreases.These layers aim to extract features of image.Additionally, the researchers use max pooling layers in multiple stages for downsampling which helps model learn more features in different levels.
Decoder
The purpose of decoder is to generate a segmented mask which has the same dimension as input image.In order to meet that requirement, using Upsampling process combined with convolutional layers in between. This technique helps reconstruct fine-grained features,thus generate dense feature maps.
Input of the decoder is feature maps of encoder.Let's step back a little bit to understand how decoder uses feature maps of encoder.
After each convolutional layer of encoder,we store the feature maps.Similarly, after each pair of Upsampling layer and convolutional layer, we get other feature maps.We concatenate those two kind of feature maps ( ofcourse they must have the same dimension),meaning the model is less likely to forget what it previously learned and prone to vanishing gradient during backpropagation.
Once again, to avoid overfitting, the researchers employed Regularization technique with L2 norm.
Calibration
In the paper, the authors proposed a metric called confidence to help models avoid ill-calibrated which makes them overconfident about their predictions.The output of XNet is a 3-dimensional probability map where each pixel is assigned a probability of belonging to one of the three different categories we mentioned previously.
Network confidence is defined as:
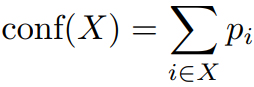
X is a set of pixels assigned to a category(e.g.bone,soft tissue), and p_i is the probability that pixel ith in X belongs to said category.
False Positive Reduction
The authors want to reduce the soft tissue false positives, meaning that if model classifies a pixel as soft tissue, we want the probability of that case higher and less likely to be wrong.That decision making process requires comparison of that probability with a given threshold.Therefore,in order to meet that demand, we increase the threshold at the expense of the quantity of true positives.As you might realize, this is actually a trade-off process.
Result
Because the authors were looking for a balance between false positive reduction and true positive enhancement,"Accuracy" might not be a suitable metric.Instead, they chose F1 score to measure network performance.
Actually, they still compared performance of network in different metrics.The result was summarized in the table 3.
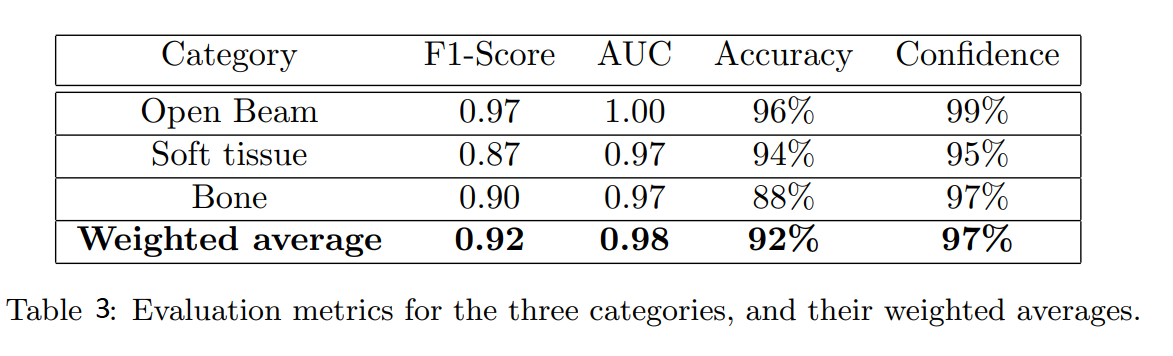
Conclusion
XNet was designed as an end-to-end model which has some advantages:
- Gains high accuray and outperforms other networks on a small dataset.This helps medical institution reduce the cost of labelling,only needs a little data but still get a good result.
- It was well-calibrated,evaluated on different metrics,avoiding bias and overconfident.
- Its architecture is easy to implement.
With those pros, XNet has high potential to apply widely.
