
Open-Source Internship opportunity by OpenGenus for programmers. Apply now.
Reading time: 30 minutes
A clique is a subset of vertices of an undirected graph G such that every two distinct vertices in the clique are adjacent; that is, its induced subgraph is complete. Cliques are one of the basic concepts of graph theory and are used in many other mathematical problems and constructions on graphs.
The task of finding whether there is a clique of a given size in a graph (the clique problem) is NP-complete, but despite this hardness result, many algorithms for finding cliques have been studied.
A maximal clique is a clique that cannot be extended by including one more adjacent vertex, that is, a clique which does not exist exclusively within the vertex set of a larger clique.
A maximum clique of a graph, G, is a clique, such that there is no clique with more vertices. Moreover, the clique number ω(G) of a graph G is the number of vertices in a maximum clique in G.
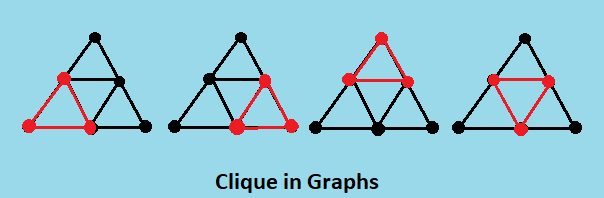
Theorems regarding Clique
-
Turán's theorem gives a lower bound on the size of a clique in dense graphs. If a graph has sufficiently many edges, it must contain a large clique. For instance, every graph with n vertices and more than ⌊ n⁄2 ⌋ ⋅ ⌈ n⁄2 ⌉ edges must contain a three-vertex clique.
-
Ramsey's theorem states that every graph or its complement graph contains a clique with at least a logarithmic number of vertices.
-
According to a result of Moon & Moser (1965), a graph with 3n vertices can have at most 3n maximal cliques. The graphs meeting this bound are the Moon–Moser graphs.
Classes of graphs may be characterized by their cliques:
- A cluster graph is a graph whose connected components are cliques.
- A block graph is a graph whose biconnected components are cliques.
- A chordal graph is a graph whose vertices can be ordered into a perfect elimination ordering, an ordering such that the neighbors of each vertex v that come later than v in the ordering form a clique.
- A cograph is a graph all of whose induced subgraphs have the property that any maximal clique intersects any maximal independent set in a single vertex.
- An interval graph is a graph whose maximal cliques can be ordered in such a way that, for each vertex v, the cliques containing v are consecutive in the ordering.
- A line graph is a graph whose edges can be covered by edge-disjoint cliques in such a way that each vertex belongs to exactly two of the cliques in the cover.
- A perfect graph is a graph in which the clique number equals the chromatic number in every induced subgraph.
- A split graph is a graph in which some clique contains at least one endpoint of every edge.
- A triangle-free graph is a graph that has no cliques other than its vertices and edges.
What is the Clique Problem?
The clique problem is the computational problem of finding cliques in a graph. It has several different formulations depending on which cliques, and what information about the cliques, should be found.
Common formulations of the clique problem include finding a maximum clique, finding a maximum weight clique in a weighted graph, listing all maximal cliques, and solving the decision problem of testing whether a graph contains a clique larger than a given size.
Most versions of the clique problem are hard. The clique decision problem is NP-complete (one of Karp's 21 NP-complete problems). The problem of finding the maximum clique is both fixed-parameter intractable and hard to approximate. And, listing all maximal cliques may require exponential time as there exist graphs with exponentially many maximal cliques.
Is Clique decision problem NP-complete?
A problem is NP-complete when it can be solved by a restricted class of brute force search algorithms and it can be used to simulate any other problem with a similar algorithm. More precisely, each input to the problem should be associated with a set of solutions of polynomial length, whose validity can be tested quickly (in polynomial time), such that the output for any input is "yes" if the solution set is non-empty and "no" if it is empty. The complexity class of problems of this form is called NP, an abbreviation for "nondeterministic polynomial time". A problem is said to be NP-hard if everything in NP can be transformed in polynomial time into it, and a problem is NP-complete if it is both in NP and NP-hard. The NP-complete problems represent the hardest problems in NP.
The clique decision problem is NP-complete. It was one of Richard Karp's original 21 problems shown NP-complete in his 1972 paper "Reducibility Among Combinatorial Problems". Because of the hardness of the decision problem, the problem of finding a maximum clique is also NP-hard. If one could solve it, one could also solve the decision problem, by comparing the size of the maximum clique to the size parameter given as input in the decision problem.
Some NP-complete problems (such as the travelling salesman problem in planar graphs) may be solved in time that is exponential in a sublinear function of the input size parameter n, significantly faster than a brute-force search. However, it is unlikely that such a subexponential time bound is possible for the clique problem in arbitrary graphs, as it would imply similarly subexponential bounds for many other standard NP-complete problems.
Algorithms/ Problems
Finding a single maximal clique
A single maximal clique can be found by a straightforward greedy algorithm. Starting with an arbitrary clique (for instance, any single vertex or even the empty set), grow the current clique one vertex at a time by looping through the graph's remaining vertices. For each vertex v that this loop examines, add v to the clique if it is adjacent to every vertex that is already in the clique, and discard v otherwise. This algorithm runs in linear time. Because of the ease of finding maximal cliques, and their potential small size, more attention has been given to the much harder algorithmic problem of finding a maximum or otherwise large clique than has been given to the problem of finding a single maximal clique.
Cliques of fixed size
One can test whether a graph G contains a k-vertex clique, and find any such clique that it contains, using a brute force algorithm. This algorithm examines each subgraph with k vertices and checks to see whether it forms a clique. It takes time O(nkk2), as expressed using big O notation. This is because there are O(nk) subgraphs to check, each of which has O(k2) edges whose presence in G needs to be checked. Thus, the problem may be solved in polynomial time whenever k is a fixed constant. However, when k does not have a fixed value, but instead may vary as part of the input to the problem, the time is exponential.
Listing all maximal cliques
To find a maximum clique, one can systematically inspect all subsets, but this sort of brute-force search is too time-consuming to be practical for networks comprising more than a few dozen vertices. Although no polynomial time algorithm is known for this problem, more efficient algorithms than the brute-force search are known. For instance, the Bron–Kerbosch algorithm can be used to list all maximal cliques in worst-case optimal time, and it is also possible to list them in polynomial time per clique. By a result of Moon & Moser (1965), any n-vertex graph has at most 3n⁄3 maximal cliques, and the worst-case running time of the Bron–Kerbosch algorithm (with a pivot strategy that minimizes the number of recursive calls made at each step) is O(3n⁄3), matching this bound.
What are the applications of Clique?
-
Let's consider a social network, where the graph's vertices represent people, and the graph's edges represent mutual acquaintance. Then a clique represents a subset of people who all know each other, and algorithms for finding cliques can be used to discover these groups of mutual friends.
-
The clique problem has many applications in bioinformatics. For instance, Ben-Dor, Shamir & Yakhini (1999) model the problem of clustering gene expression data as one of finding the minimum number of changes needed to transform a graph describing the data into a graph formed as the disjoint union of cliques.
-
In electrical engineering, Prihar (1956) uses cliques to analyze communications networks
-
Paull & Unger (1959) use them to design efficient circuits for computing partially specified Boolean functions.
-
The clique problem also has many applications in computational chemistry. For instance, Kuhl, Crippen & Friesen (1983) use cliques to model the positions in which two chemicals will bind to each other.
Related articles
Using Bron Kerbosch algorithm to find maximal cliques in O(3^(N/3))Greedy approach to find a single maximal clique in O(V^2) time complexity
Algorithm to find cliques of a given size k 【O(n^k) time complexity】
